Lab Technology
Targeted mass spectrometry; live-cell fluorescence microscopy; high-throughput screening of chemical, RNAi and DNA construct libraries; and computational modeling are among the tools we use for research.
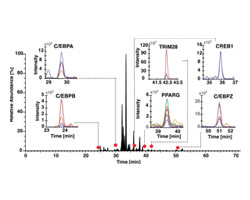
QUANTITATIVE TARGETED PROTEOMICS
We are using Selective Reaction Mass Spectrometry to obtain quantitative protein profiles of signaling networks. Our lab has a Thermo TSQ Vantage triple-quadropole mass spectrometer with Proxeon NanoHPLC dedicated to this approach.
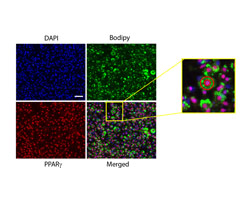
SINGLE-CELL IMAGING
We use live and fixed microscopy to understand the single-cell dynamics underlying cell decision processes
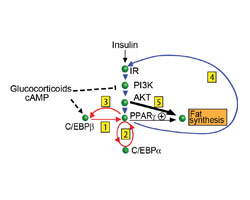
MODELING
Static diagrams cannot capture the dynamic interactions of signaling networks. We use quantitative modeling to be able to make sense of networks in time and space.

